One of the common problems with Sanger sequencing is samples with high salt concentration. In general, one could talk about inhibitors as such, but salts are probably the most common case. If you send us such samples, you will probably not be very satisfied with the quality of the result. In this article, we will take a closer look at why this is the case and, most importantly, what can be done about it.
Why salts are a problem
Sanger sequencing is basically nothing more than an enzymatic reaction plus electrophoresis. The reaction takes place using commercial mixes and any ionic imbalance will of course have a negative effect on its course.
Electrophoresis in automated analyzers is then performed in capillaries. If there are unwanted impurities in the sample, then of course it depends on how they behave in the electric field. Salt ions are a big problem. These are charged particles, so they enter the capillaries, and because they are small, they migrate through them very quickly, much like small DNA fragments.
Thus, any salts in the sent template may cause problems at different stages of its processing and may affect results in different ways.
As far as the sequencing reaction itself is concerned, in this step, the result cannot be influenced in any way on our side during routine processing. We take the template as you give it to us, including salinity, we do not clean it before the reaction, so we do not remove any inhibitors. Subsequent treatment of the sequencing reaction products is performed so that (among other things) all salts are removed before the actual electrophoresis. This means that after the sequencing reaction, its products must be purified, and one of the reasons for doing so is desalination. Pure DNA then dissolves in some deionized matrix from which it is "injected" into the capillary.
Thus, the ideal situation is that the sequencing reaction is performed on pure DNA in a reaction mixture which, of course, contains a small amount of salt (as part of the reaction buffer), but these are subsequently removed in the reaction purification step. The obtained pure product enters the capillaries from the deionized matrix and its size separation takes place, which results in beautiful data.
What happens if there are salts in the sample
Of course, we see the salts in the sample only after electrophoresis, before that this is not technically possible. Typically, the salts in the capillary appear as a white area at the beginning of the electropherogram, but you must look mainly at the raw data. It looks something like this, the above salty sample, the bottom clean, the salts are the white area at the beginning of the raw data (see ellipse):
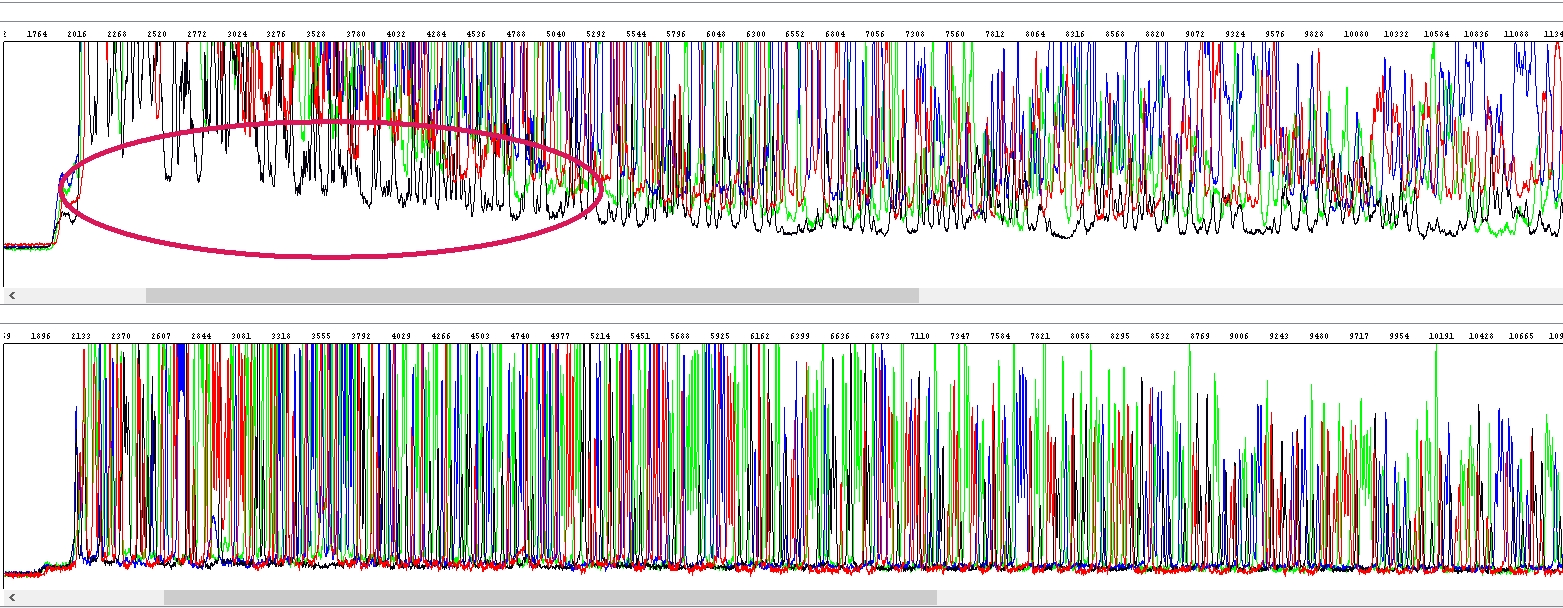
Notice the increased background precisely because of their presence. Depending on the salt content, this "white area" can be larger or smaller and this of course has an effect on what the data in the electropherogram look like:
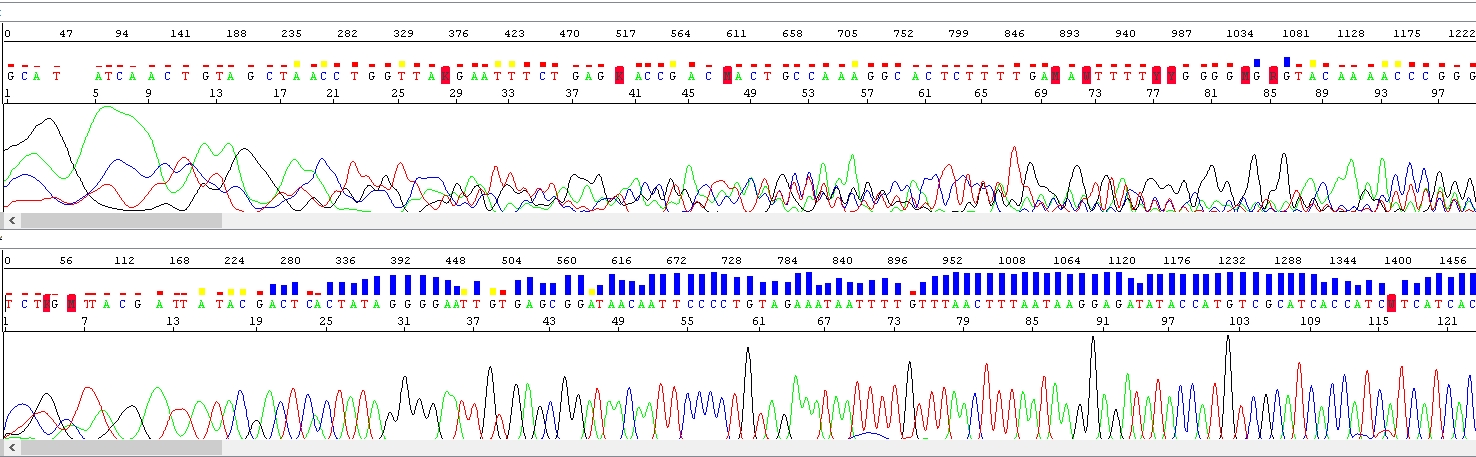
This is an example of an extremely salty sample! At first glance, the electropherogram above looks like a mixture, but the cause is elsewhere, purification of the sequencing reaction did not remove salt.
If you get such a result from us, there is nothing you or we can do about it. By repeating the electrophoresis we get the same picture. The sample could theoretically be repurified in an effort to remove those salts, but in practice, each purification is associated with DNA loss and we definitely do not recommend or offer it. Honestly, it's not worth the effort. First of all, it is necessary to learn from this and change the working procedure for the preparation of templates so that this situation does not occur next time. So at this point, the question arises, where did the salts come from and what should I change?
Where can salts come from?
We basically sequence two types of products, plasmids and PCR products. The ideal situation is that both types of templates are always cleaned by you (among other things, desalted) and then a sequencing reaction is performed on them. In practice, however, it often happens that the client facilitates the work or some commercial kits do not work optimally. An example is the so-called colony PCR. Although the PCR reaction will work, there are actually various contaminations in the test tube after it has been performed. If you subsequently send us such a sample, the result may or may not be satisfactory, again depending on the content of contaminants. Their presence can inhibit the course of the sequencing reaction itself (not to mention the electrophoresis), so its yield will be low as well as signals, despite the fact that, of course, each sequencing reaction is always purified by us.
Even if you purify such a PCR product, it may not be enough. Some methods of purification of eg PCR products are more suitable than others and it is always necessary to keep in mind that what applies to one may not apply to the other. A very good and efficient way to purify PCR products is ExoSAP. It is a simple and robust method that ensures nucleotide dephosphorylation and primer degradation, but never removes any salts or other contaminants. So if you have a problem with the high salt content in the templates for your PCR, purifying PCR products with ExoSAP is not exactly the best you can choose, although the method is very suitable for other PCR products.
Similarly, for example, column cleaning can turn into a nightmare, because despite decades of column kit development, some are still annoying, see the older article here.
In summary, it pays to know the principle of the different procedures available and not just rely on the various commercial instructions for the kits, the composition of which, in most cases, you do not know at all.
Examples of results
If you have a problem with the salt content in the samples, the results may look different. You can see one example in the picture above and below are a few more examples.
Example no. 1:
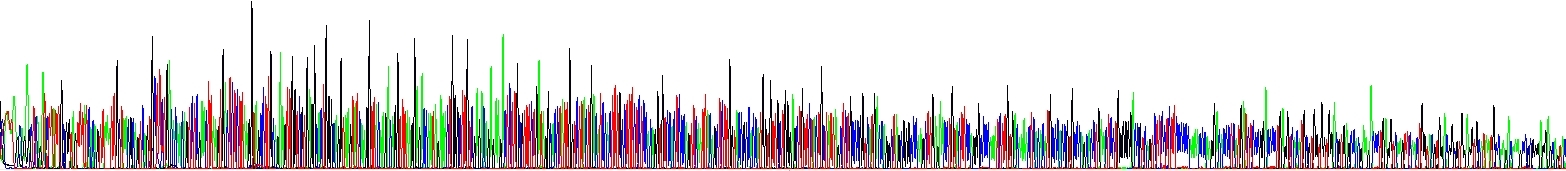
Correct salt concentration
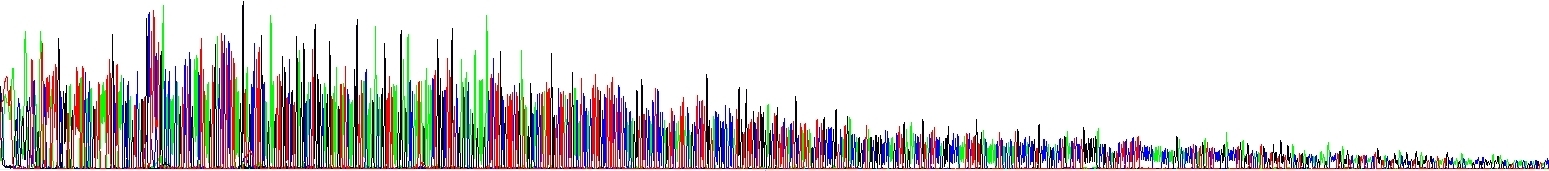
The sequencing reaction was performed with the same template and in the same manner, but NaCl was added to the second sample. The amount of salts in the sample is therefore increased, but only slightly (compare with the picture above). Notice the falling signal. This is a typical result when the salt concentration is increased but still not too high and there is no complete inhibition of the sequencing reaction. At the same time, the salts were effectively removed before the electrophoresis itself, the background is not increased.
Example no. 2:
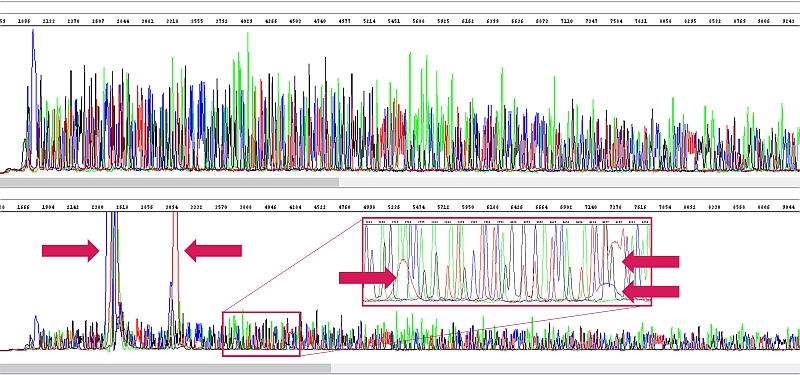
There was a high content of some inhibitor in the samples (in this case, unfortunately, we are not sure that it was really a salt). As a result of its presence, not only did the signal intensity and thus the read length decrease, but so-called blobs (see arrows) also occurred in the reaction, which make the interpretation of sequencing data problematic wherever they occur. Since the DNA concentration in these samples was good, we diluted the samples three times at the client's request and repeated the reaction (of course, the template must be diluted in the sequencing reaction, not the product into the capillary). This result is already OK (above sequencing sequencing reaction after template dilution, below original sample with blobs).
Note on blobs - These are residual dye terminators. In reality, the DNA concentration may either be low or good, but inhibitors are present at the same time. Thus, the sequencing reaction proceeds in low yield, which corresponds to the low consumption of terminators. Subsequent purification of the sequencing reaction fails to remove all these free terminators and you can see them in the electropherogram. It means: little DNA or inhibition = low reaction yield = low signal in the electropherogram and sometimes blobs = mostly short read length and worse data quality at the beginning (due to salts) and in the blob area. Therefore, if there is little DNA in the reaction, more must be added next time. However, if this condition is caused by DNA contamination and thus inhibition of the sequencing reaction, it may help to use less DNA (but not little) or it needs to be better purified.
Example no. 3:
Sometimes the situation is more complicated. For example, if contamination (salts) occurs in combination with some other problem, the result can be confusing and interpretation problematic, especially if you do not have enough experience with it. Below is an example of a poorly purified and also mixed sample:
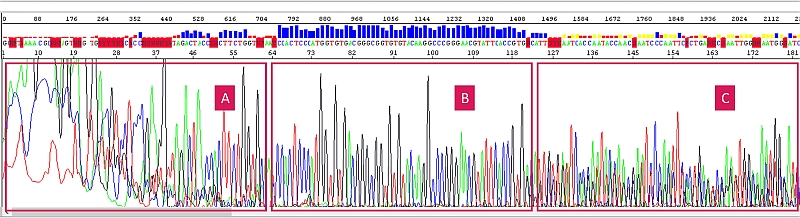
A – the sequence is harder to read due to salinity, B – the sequence is fine, the salts have already migrated away, C – the sequence is salt-free but mixed (mixed template).
Of course, other factors can come into play, such as signal strength, etc. So there are many possibilities for what the results may look like. Take the above examples as illustrative, not a complete list of options.
Short summary
In short, it is not surprising that if you want to have good results, you need to supply clean DNA. If your results look like the ones above, you need to analyze why this is the case and choose the appropriate corrective action, which this article will hopefully help a bit. Unfortunately, due to the routine nature of the work, there is almost no room for optimization on our side. Theoretically, we could, for example, dilute samples, etc., but in practice this always means a charged repetition of the analysis.
Keep in mind that:
In conclusion, we would like to point out that all recommendations must be taken as indicative and it is always necessary to correctly assess your specific situation. If you have any questions, do not hesitate to contact us.
Note: We also recently published an article describing a comparison of different purification kits. It is quite possible that the low quality of the result of some of them (and, for example, the short read length) is to some extent (possibly exclusively) caused by the inhibitors contained in the product of the purification.
Sanger lab, info@seqme.eu